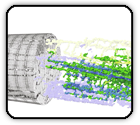
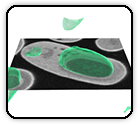
Semi-Automated 3D segmentation of human skeletal muscle using Focused Ion Beam-Scanning Electron Microscopic images
Caffrey BJ, Maltsev AV, Gonzalez-Freire M, Hartnell LM, Ferrucci L, Subramaniam S. Journal of Structural Biology 207: 1-11 (2019)
The cryo-EM revolution: fueling the next phase
Subramaniam S. IUCrJ 6: 1-2 (2019)
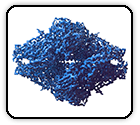
Single-particle cryo-EM structure of a voltage-activated potassium channel in lipid nanodiscs
Matthies D, Bae C, Toombes G, Fox T, Bartesaghi A, Subramaniam S, Swartz K. eLife 7: e37558 (2018)
Atomic Resolution Cryo-EM Structure of β-Galactosidase
Bartesaghi, A, Aguerrebere, C, Falconieri, V, Banerjee, S, Earl, LA, Zhu, X, Grigorieff N, Milne, JLS, Sapiro, G, Wu, X, Subramaniam, S.
Structure 26(6), 848-856. (2018)
Microbiology catches the cryo-EM bug
Earl L, Falconieri V, Subramaniam S. Current Opinion in Microbiology 43: 199-207 (2018)
Cryo-EM structure of human rhodopsin bound to an inhibitory G protein
Kang, Y, Kuybeda, O, de Waal, PW, Mukherjee, S, Van Eps, N, Dutka, P, Zhou, XE, Bartesaghi, A, Erramilli, S, Morizumi, T, Gu, X, Yin, Y, Liu, P, Jiang, Y, Meng, X, Zhao, G, Melcher, K, Ernst, OP, Kossiakoff, AA, Subramaniam, S, Xu, HE.
Nature 558(7711), 553-558. (2018)
Deep-learning-assisted Volume Visualization
Cheng H-C, Cardone A, Jain S, Krokos E, Narayan K, Subramaniam S, Varshney A. IEEE Transactions on Visualization and Computer Graphics 25: 1378-91 (2018)
Structural mechanisms of centromeric nucleosome recognition by the kinetochore protein CENP-N
Chittori S, Hong J, Saunders H, Feng H, Ghirlando R, Kelly AE, Bai Y, Subramaniam S. Science 359: 339-43 (2018)
Cryo-EM noted as one of the biggest scientific breakthroughs of 2017
Dr. Subramaniam’s work on resolving the structures of beta-galactosidase and nucleosomes (DNA-Protein complexes) was featured in Science Magazine’s Breakthroughs of the year video, featuring cryo-electron microscopy (cryo-EM) among the biggest scientific breakthroughs of 2017.
Cryo-EM structures reveal mechanism and inhibition of DNA targeting by a CRISPR-Cas surveillance complex
Guo TW, Bartesaghi A, Yang H, Falconieri V, Rao P, Merk A, Eng ET, Raczkowski AM, Fox T, Earl LA, Patel D, Subramaniam S. Cell 171: 414-26 (2017)